|
Detected gene hubs and hub sub-networks were integrated by signature loci associated with cancer that include, for example, NOTCH1, RAC1, PIK3CD, BCL2, and EGFR. This integrative network analysis approach revealed that gene expression and methylation consistently targeted the same gene pathways relevant to cancer: Pathways in cancer, Ras signaling pathway, PI3K-Akt signaling pathway, and Rap1 signaling pathway, among others. The application of a novel signal detection-machine learning approach to methylation analysis of whole genome bisulfite sequencing (WGBS) data permitted a high level of methylation signal resolution in cancer-associated genes and pathways. With the purpose of identifying reliable cancer-associated methylation signal in gene regions from leukemia patients, we present an integrative network analysis of differentially methylated (DMGs) and differentially expressed genes (DEGs). 
Integrated network analysis of cytosine methylation and expression datasets has the potential to provide deeper insights into the complex disease states and their causes than individual disconnected analyses. 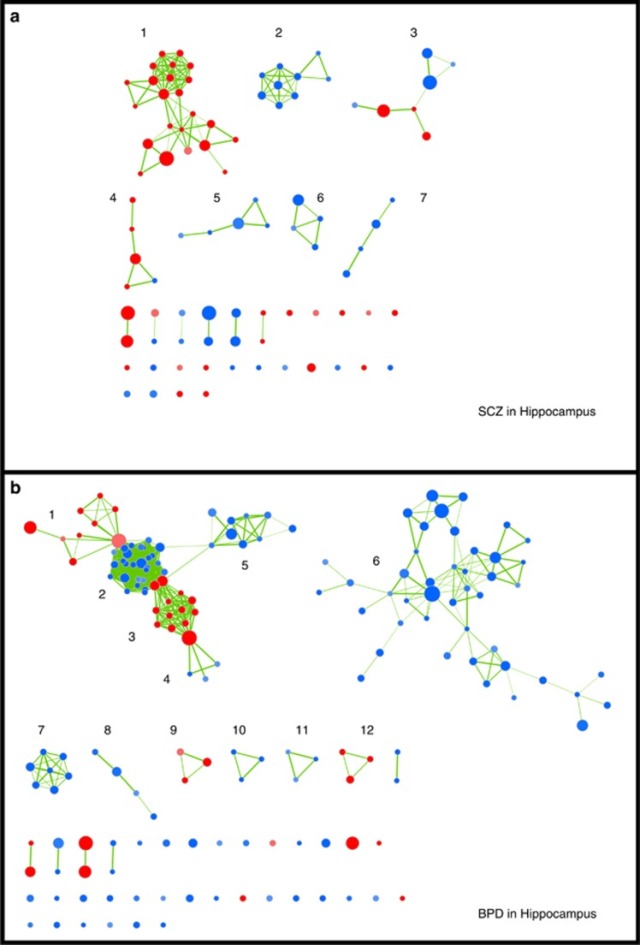 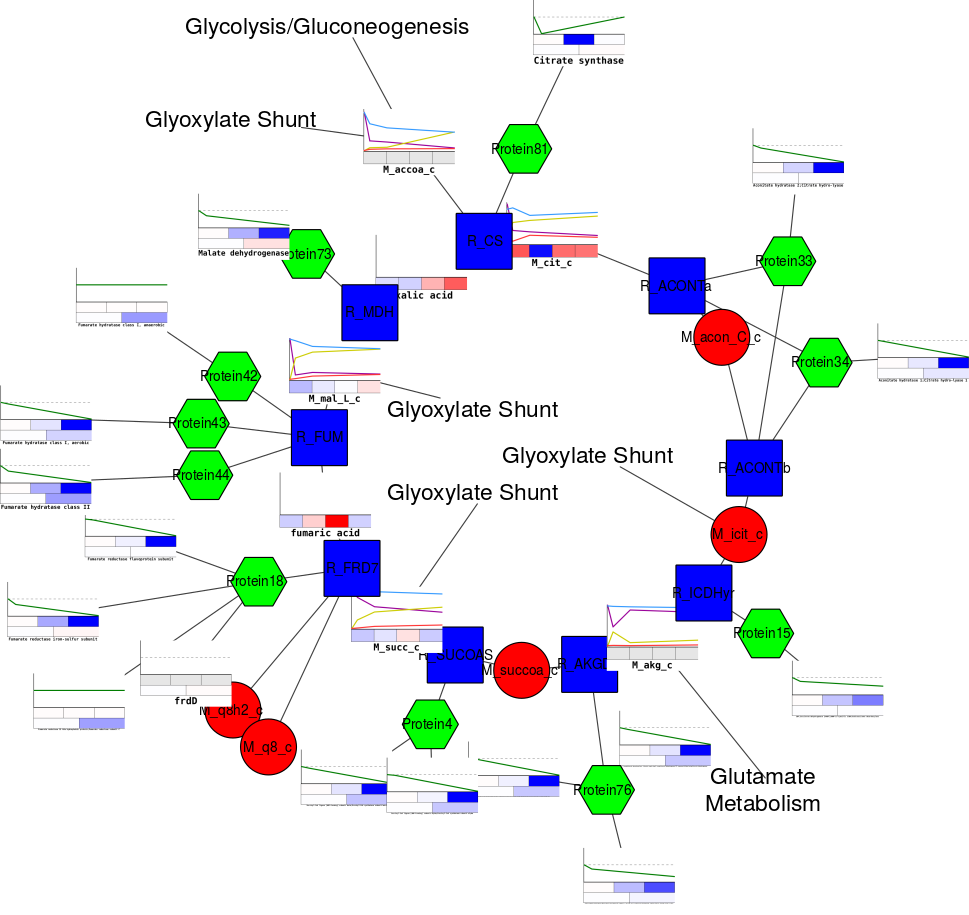
Genome-wide DNA methylation and gene expression are commonly altered in pediatric acute lymphoblastic leukemia (PALL).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
